Points of View from CERN Alumni: Synthetic Biology
I think the biggest innovations of the 21st century will be at the intersection of biology and technology. A new era is beginning.
--Steve Jobs
Steve Jobs did not live to see his vision become reality. But today, biology is quickly becoming a quantitative discipline where physicists and engineers will feel right at home. It will give rise to some of this century’s greatest commercial and humanitarian opportunities, as well as its most profound ethical controversies.
This article gives a brief overview of the emerging field of synthetic biology. If you are contemplating life after CERN, or if you are curious about where the world is headed, synthetic biology should be on your radar screen. If you would like to know more about it, please drop me a note. I am always happy to chat with CERN alumni!
All life uses the same operating system
All living things, from bacteria to plants to people, use the same system of genetic codes based on the familiar A, T, C and G nucleotides. In a sense, living things are like computers that all use the same operating system. The quest to understand how genetic codes impact living systems is at the heart of synthetic biology.
Synthetic biology (or synbio for short) is an engineering approach to biology, aimed at designing and fabricating biological systems that do not already exist in the natural world. It harnesses the power of biology in unprecedented ways for purposes such as producing therapeutic drugs or converting biomass to chemicals. Few areas of our lives will be untouched by it.
Synbio is already being used to design personalized medicines that are more effective than existing alternatives, with fewer side effects. It is also creating diagnostics that identify diseases in real time without time-consuming lab tests, potentially averting epidemics. And in the years ahead, it will allow researchers to develop reasonably priced treatments for rare diseases.
In agriculture, synthetic biology is redefining what we grow and how we grow it. Crops can be engineered for bacteria resistance, for example, or enhanced photosynthesis or reduced spoilage.
Synbio promises to give a big boost to the environment as well, through innovations such as nitrogen-fixing bacteria that reduce the need for environmentally harmful fertilizers. Its early success stories include bacteria that produce high-value petrochemicals without using petroleum. And last year, researchers developed an enhanced enzyme that breaks down the PET (polyethylene terephthalate) plastic used for soft drink bottles, recycling it into its original components.
Synbio will even disrupt fields that we do not currently think of as biological. Living systems produce the most complex structures in the universe with remarkable efficiency. With this in mind, the Defense Advanced Research Projects Agency (DARPA) in 2013 launched a synthetic biology initiative to create 1000 new materials by means of metabolic processes. This approach could vastly expand the scope of chemistry and materials science.
Why didn’t anyone do synthetic biology before?
The idea of using engineered organisms as a means of production has been around since 1978, when Genentech started manufacturing synthetic human insulin using genetically modified E. coli bacteria. But only recently has synthetic biology developed into a flexible platform to create special-purpose biological systems. This newfound capability is due to rapid advances in the technology to read, write and edit DNA, plus progress in robotics and machine learning.
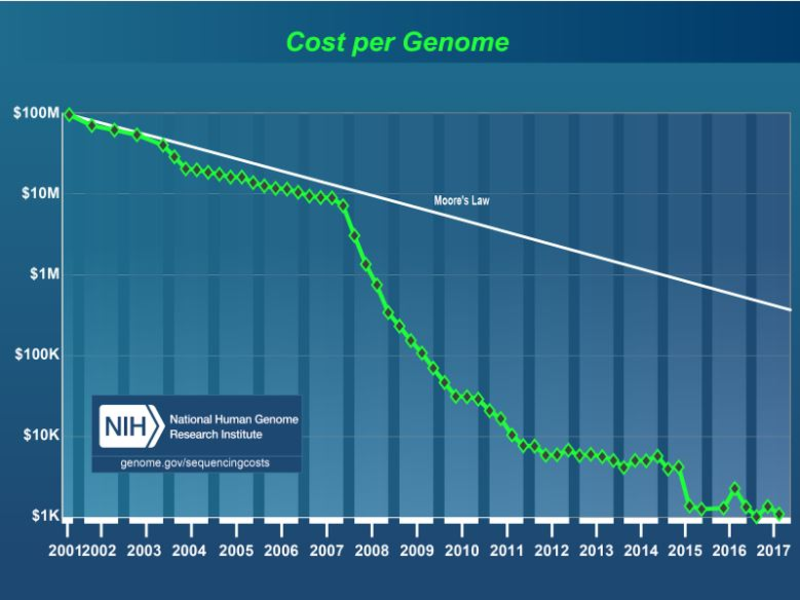
In 2003, the Human Genome Project and Celera Corporation successfully mapped all 3 billion base pairs of the human genome. Since then, the price of gene sequencing has fallen by more than 50% per year, much faster than Moore’s Law (Figure 1). Today a comparable genome could be sequenced for about $1000. The cost of writing DNA is plummeting too, though it still costs far more to write DNA than to read it. Easy access to low-cost synthetic DNA will be a driving force behind the next wave of innovation in synthetic biology.
Without question though, the most publicized tool of synthetic biology is gene editing, especially CRISPR. CRISPR has grabbed headlines with its revolutionary ability to make precise edits to existing genomes. This has raised urgent concerns on issues ranging from designer babies to potentially irreversible changes to the biosphere. Meanwhile, CRISPR has become a staple of biological research. It is a simple way to make multiple edits to genomes, either to deactivate code segments or to replace them.
Figure 1
The dawn of digital biology
The nature of biological lab work has not changed much in the last hundred years. Outside of pharmaceutical manufacturing, biology experiments are still typically done by individual researchers working by hand, one step at a time. Measurements are often imprecise, results are difficult to share, and experiments are difficult — if not impossible — to reproduce. The newest generation of lab equipment addresses these shortcomings though, with big implications for how biology is done, both commercially and in academia.
Inexpensive robotic lab equipment, with open-source APIs, allows researchers to transfer their results straight from the laboratory to the production facility. Instead of using handwritten notebooks, biologists and engineers can now store experimental protocols in code repositories, like a Github of biological manufacturing processes. Just as importantly, robotic equipment allows researchers to do multiple experiments simultaneously. The settings of all lab instruments, plus all the genetic and physiological features of engineered organisms, are now parameters that can be rapidly optimized in parallel.
As the quantity and quality of experimental data improve, biology is becoming increasingly data-driven. Synthetic biologists are moving away from the scientific method of hypothesis testing. Instead, they use discovery science and algorithmic experimentation. Discovery science stresses empirical analysis of large volumes of experimental data to find new patterns or correlations. Algorithmic experimentation, also known as active learning, is a subfield of machine learning focused on algorithms that select the data they need to improve their own models. Together, discovery science and algorithmic experimentation turbocharge evolution to design biological systems for specific purposes.
The age of lean biotech startups
Since its inception a generation ago, the field of synthetic biology has mostly been government-funded, primarily through DARPA and the National Science Foundation. It has focused on basic research to develop an engineering toolset for future investigations and product development. Until recently, there was not much of a startup scene. Even in the private sector, the large research teams and expensive equipment necessary to design new drugs or biochemicals meant that only big players in big markets could compete.
But as the core technologies become broadly available at reasonable cost, the barriers to entry are falling. As a result, biology is becoming increasingly innovative. For small startups, access to low-cost plug-and-play tools means they can create products that once seemed beyond their reach. Venture capitalists are responding with an influx of new investment (Figure 2).
Figure 2
Conclusion
Synthetic biology is spawning innovative startups at the intersection of biology and technology, ushering in the new era that Steve Jobs envisioned. These startups show that biotechnology is no longer the exclusive province of giant pharmaceutical companies. And because synthetic biology is inherently data-driven, they will benefit disproportionately from advances in related fields such as computing and machine learning. Synthetic biology is going to create remarkable career opportunities for physicists, offering unprecedented potential to elevate the quality of human life.
John Murray, February 2019
